 This will be used to announce new versions and advise users about bugs and problems. 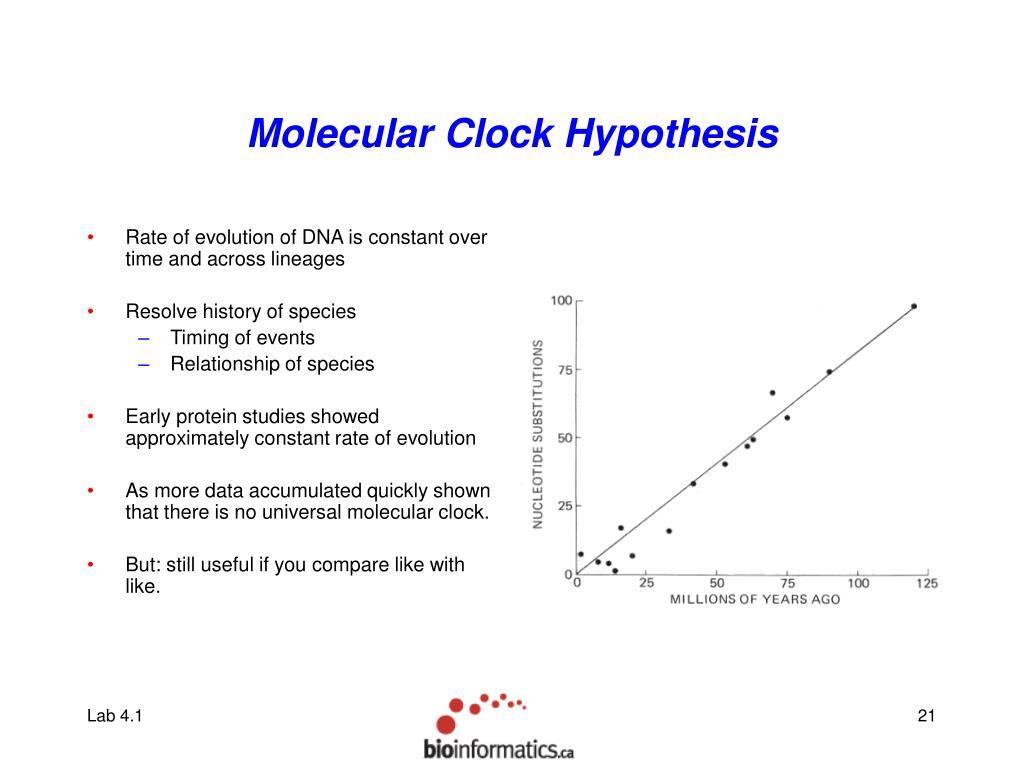
Users are strongly advised to join the BEAST mailing-list. Bioinformatics, 18, 1404-1405.īEAST is built on a large body of prior work and appropriate citations for individual modules, models and components will be listed when BEAST is run. Pybus OG & Rambaut A (2002) GENIE: estimating demographic history from molecular phylogenies. Rambaut A (2000) Estimating the rate of molecular evolution: incorporating non-contemporaneous sequences into maximum likelihood phylogenies. DOI:10.1093/ve/vey016 BEAST is descended from earlier work:ĭrummond AJ, Nicholls GK, Rodrigo AG & Solomon W (2002) Estimating mutation parameters, population history and genealogy simultaneously from temporally spaced sequence data. Citing BEAST The recommended citation for this program is: Suchard MA, Lemey P, Baele G, Ayres DL, Drummond AJ & Rambaut A (2018) Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10 Virus Evolution 4, vey016. Install BEAST on UNIX/Linux or Mac command-lineĪs an introduction to using BEAST we provide some basic introductory tutorials using the graphical applications of BEAST to perform analyses using provided example files.What can BEAST do? Getting started with BEAST Downloading BEAST For details about BEAST2, an independent project led by the University of Auckland, please look here. 
This website is for BEAST v1.X (currently version v1.10.4). We include a simple to use user-interface program for setting up standard analyses and a suit of programs for analysing the results. BEAST uses MCMC to average over tree space, so that each tree is weighted proportional to its posterior probability. It can be used as a method of reconstructing phylogenies but is also a framework for testing evolutionary hypotheses without conditioning on a single tree topology. It is entirely orientated towards rooted, time-measured phylogenies inferred using strict or relaxed molecular clock models. Phylogeographic Diffusion in Continuous Space, WNVīEAST is a cross-platform program for Bayesian analysis of molecular sequences using MCMC.3 Beyond the strict molecular clock There are several ways to deal with violations of molecular clock assumptions: 1.Use strict clock but lter the data Gene selection Lineage selection Note, that both approaches reduce the amount of data (increase variance). Phylogeographic Diffusion in Continuous Space, YFV the molecular clock is a good approximation of reality.Phylogeographic Diffusion in Discrete Space.Phylodynamics inference of respiratory viruses. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
